Alternative explanation on marker validation
Marker validation is the determination whether or not markers can predict a certain trait in the wider breeding material.
Markers are developed in mapping populations. However, a marker that is tightly linked to a gene of interest is not necessarily perfect in predicting a trait in breeding material. For example, when the genes affecting a trait are located on different parts of the genome and a specific marker is only tracking one of the genes on one chromosome, while in other material a different gene is segregating for the same trait, on a different chromosome is responsible. In other breeding lines, the genes for that trait will be associated with different markers than the one that was discovered in the mapping population.
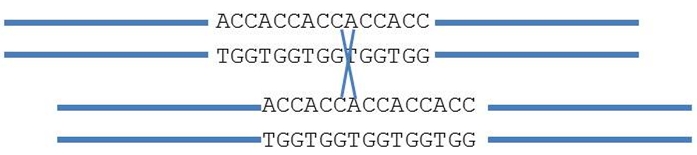
Also, marker alleles might look the same, but are in fact different. The following SSR (simple sequence repeat) marker, consists of five ACC repeats. The number of repeats determines the size of the amplification fragment. During normal pairing, fragment length does not change. However, sometimes pairing is unequal and shifted (see figure). Recombination (cross) causes the number of repeats to change to four or six ACC motifs: new alleles are formed. This may happen on several occasions resulting in alleles that appear identical (e.g. with 6 repeats) but are formed during different events.
Therefore, even markers that are very tightly linked in a mapping population have to be validated in the relevant breeding material.