Marker-assisted gene pyramiding
Gene pyramiding or gene stacking is the process of combining multiple genes for one trait into a single genotype. The most widespread application for pyramiding has been for combining multiple disease resistance genes to a same pathogen species in order to develop durable disease resistance. When several resistance genes are combined, the probability that this resistance is broken is small, since the pathogen should undergo simultaneous mutations or recombinations for the corresponding genes for avirulence. Because each resistance gene on its own provides complete resistance, it is difficult to test phenotypically whether plants have either one (e.g. R1R1r2r2) or more (e.g. R1R1R2R2) effective resistance genes. In that case, MAS may be of help.
Without MAS, for example, to discriminate the desirable genotype R1R1R2R2 (resistant) from genotypes R1R1r2r2 and r1r1R2R2 (also resistant), one needs to do several testcrosses:
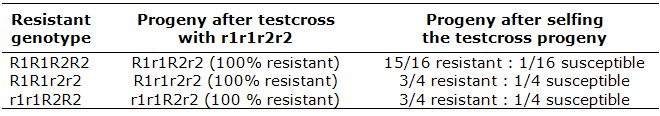
With MAS, using reliable markers, plants can be selected directly in the F2:
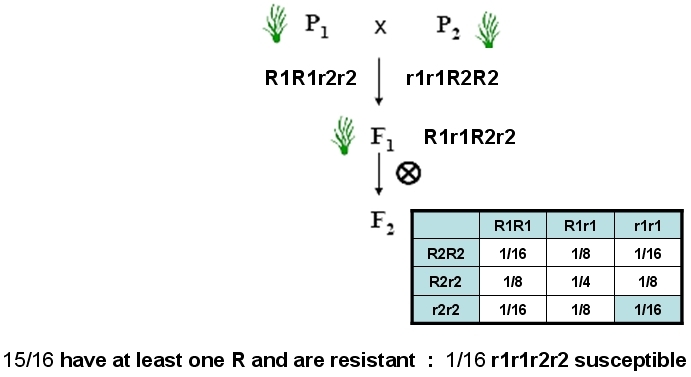
Marker-assisted pyramiding of two dominant disease resistance genes for the same pathogen.
Each resistance gene provides 100% resistance against the pathogen. Combining two different resistance genes in a variety is thought to make this resistance more durable, because the probability of simultaneously overcoming both types of resistances presumably is small. During conventional phenotypic evaluation, no difference can be observed between Rr and RR plants. Simultaneous testing of resistance genes R1 and R2 can be done using phenotypic evaluation, for example removing several leaves from the plant and inoculating the leaves individually with different isolates (one for R1 and one for R2) of the pathogen ('detached leaf assay'). However, it will remain uncertain whether plants are R1r1 or R1R1 (Adapted from B. Collard & D. Mackill (2006).)
Summary
→ MAS may be useful in gene pyramiding when each of the individual genes result in the same phenotype