Marker-assisted backcrossing (MAB)
Backcrossing implies repeated crossing with a (recurrent) parent. The recurrent parent is often a modern line or variety, with a desirable overall genotype, but lacking one or several important genes. Using conventional breeding methods, it typically takes 6–8 backcrosses to achieve a sufficiently high proportion of the recurrent parent genome. This requires a long time, during which the variety used as recurrent parent might have become outdated. The objective is to eliminate the undesirable genes in the genetic background in the first place, rather than to obtain a near-isogenic copy of a certain modern cultivar. Therefore, often not the same variety is used as recurrent parent but different modern varieties may be used successively, from BC3 or BC4 onwards.
|
Generation |
Recurrent parent genome (%) |
|
F1 BC1 BC2 BC3 BC4 BC5 BC6 |
50 75 87.8 93.75 96.88 98.44 99.22 |
Note that the expected percentage for the entire BC1 population of the recurrent parent genome is 75%, but by chance, some individuals possess more of the recurrent parent genome than 75% and others less than 75%.
Marker-assisted backcrossing may accelerate the backcrossing process by selecting against as many as the marker alleles of the donor in the background (i.e. away from the locus of the desired gene to be incorporated). In each BC generation, some plants have by chance more and others less than the statistically expected proportion of donor background. Running of the genome-wide markers. will identify individuals that happen to have a relatively low proportion of donor genotype in the background. Selection against the donor background may be done with or without a linkage map being available. See more about this subject under Principles of Plant Breeding, section Selection Methods.
- when a genetic map is available: using evenly spaced markers
- when a genetic map is not available (which is sometimes the case for exotic material): selection against all marker alleles from the donor parent, regardless of their (unknown) position in the genome.
Foreground selection (i.e. selection for the target gene to be incorporated) requires two flanking markers, to prevent selection of the donor marker, but losing the linked gene because of inadvertent recombination.
|
|
|
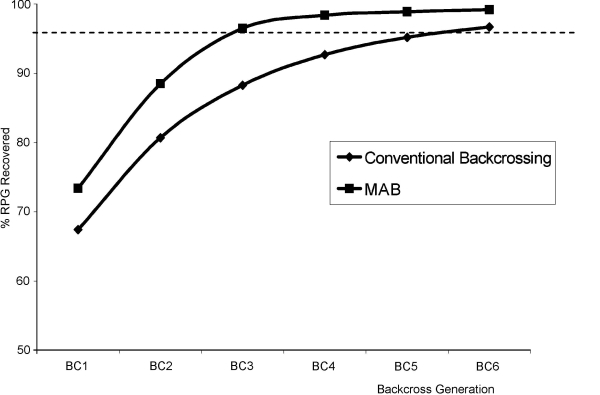
Graph of computer simulations of the percentage of recurrent parent genome (RPG recovery) in the total genome, using marker-assisted backcrossing (MAB) and conventional backcrossing. The use of markers can reduce the number of generations required to achieve the desired proportion of the recurrent parent genome (here 95%, indicated by the dotted line). (Source: Collard et al., 2005). |
(Dis)advantages of MAB
Considerable time savings can be made with MAB compared to conventional backcrossing, which could lead to economic benefits, resulting from the accelerated release of an improved variety. On the other hand, the initial cost involved in marker-assisted backcrossing would be higher than with conventional backcrossing and large investments are sometimes needed:
- Many markers are required, well spread across the genome, and preferably a genetic map, which usually is not available for exotic donors
- Markers that are associated with a gene of interest need to be converted to diagnostic markers or haplotypes that can be run in high throughput assays
- It is not sure whether a gene discovered to be beneficial in one genetic background will also have such a beneficial effect after transfer into a different genetic background. Many genes probably perform through interactions with other genes. Genes taken out of that context may lead to disappointing results.
Summary
→ MAB can be used to minimize linkage drag (see Principles of Plant Breeding, Backcross Breeding) and to maximize recovery of the recurrent parent
Show/hide comprehension question...
Show/hide comprehension question...
I want an alternative explanation | Move on to: Marker-assisted gene pyramiding