High-resolution mapping
Why use high-resolution mapping?
Mapping studies may reveal linkage between genes and markers. However, in most cases, this linkage is not tight enough to be useful in marker-assisted breeding programs. This 'loose' linkage between a marker and a gene results in a too high percentage of recombination events between gene and markers, resulting in unreliable selection. Preferably, the mapping distance between marker and a gene has to be reduced to less than 5 cM, or rather < 1cM. This can be done by using larger mapping populations and a larger numbers of markers. This process is called high-resolution mapping.
The best marker is based on the DNA sequence of the desired allele of the gene itself. Variations in all traits are caused by variations in the DNA, which can sometimes also be made visible in markers.
|
|
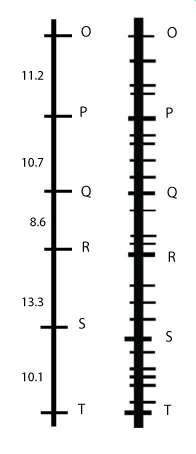
Suppose Q is the target gene of interest. Markers R and P are too far away to make them full-proof as indicator of the Q allele. Saturating the region with more markers results in markers that are more closely linked with Q than P and R were.
Note: The first picture in this module, in Markers, page 1 'What is a genetic marker', shows a gene of interest which is flanked by two markers, which are located right at the begin and the end of the gene. This was an illutration to make things clear. However, the probability of finding markers at those exact positions is very low. |
Alternatives for high-resolution mapping
If the position of a gene and two flanking markers is already known, one can use these two markers instead of high-resolution mapping. Even when the flanking markers are not very tightly linked to the gene of interest, they can be used for MAS, because double recombination within a circa 6 cM region is very rare. We discussed this in the chapter on QTL analysis in the section on Simple-interval mapping.